Introduction
Antimicrobial resistance (AMR) continues to undermine effective treatment of infectious diseases worldwide, making precise and rapid diagnostic methods essential in both clinical and research settings. Traditional culture based susceptibility testing remains a cornerstone of diagnostics, but novel tools including molecular, genomic, and rapid biochemical assays are transforming how resistance is identified and managed. This blog explores the state of the art techniques for detecting AMR, their scientific foundations, clinical utility, limitations, and future directions.
Traditional Culture-Based Antimicrobial Susceptibility Testing (AST)
1. Overview and Scientific Basis
Culture-based AST involves isolating microorganisms from patient specimens and exposing them to antimicrobial agents to determine susceptibility patterns. The approach relies on in vitro phenotypic growth inhibition, assessed via measurable endpoints such as bacterial growth zones or minimum inhibitory concentrations (MICs).
2 Key Techniques
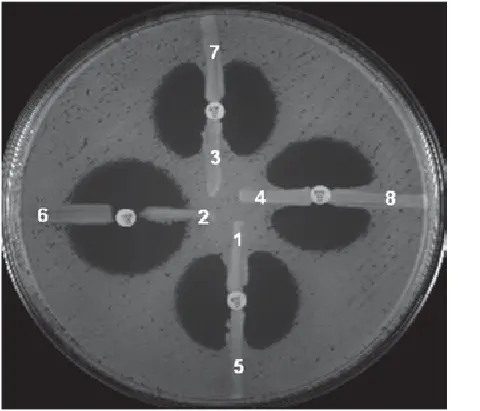
A standardized method where antimicrobial impregnated disks diffuse drugs into agar seeded with the test organism. Zones of inhibition are measured and compared to reference breakpoints. This method is recommended by clinical microbiology standards due to reproducibility and ease of interpretation.
Users can participate in beta testing programs, providing feedback on upcoming releases and influencing the future direction of the platform. By staying current with updates, you can take advantage of the latest tools and features, ensuring your business remains competitive and efficient.
3 Limitations
Time to result: 18–48 hours or longer for slow-growing organisms
Cannot directly specify resistance mechanisms
Labor-intensive in high-throughput settings
Molecular Diagnostics for AMR Detection
Polymerase Chain Reaction (PCR)-Based Assays
PCR amplifies specific genetic sequences associated with known resistance genes ( blaCTX-M, mecA, vanA). Real-time PCR (qPCR) quantifies gene copy number, enabling detection of low abundance resistance determinants directly from clinical samples.
Scientific Insight: PCR has become indispensable due to its sensitivity and specificity, especially in detecting genes not phenotypically expressed under laboratory conditions.
Multiplex PCR and Microarrays
Multiplex PCR enables simultaneous detection of multiple resistance genes in a single reaction. Microarrays expand this capacity, hybridizing amplified products to gene probes for parallel detection.

Next-Generation and Emerging Technologies
Metagenomic Sequencing
Metagenomics directly sequences all genetic material in a sample, enabling detection of resistant organisms and genes without culture. This approach is powerful for complex or polymicrobial specimens.
Scientific Application: Pendleton et al. (2017) demonstrated metagenomics in clinical AMR detection, highlighting its potential and challenges.
CRISPR-Based Diagnostics
CRISPR-Cas systems are being adapted for highly sensitive nucleic acid detection. CRISPR diagnostics can identify resistance genes with exceptional specificity and rapid turnaround.

Conclusion
The field of AMR diagnostics is rapidly evolving. Traditional culture-based methods continue to provide critical phenotypic data, while molecular and genomic tools deliver rapid, high-resolution insights into resistance mechanisms. Emerging technologies like CRISPR and metagenomic sequencing promise even greater precision and speed.
Effective AMR control relies on strategically combining diagnostic modalities, integrating results into clinical decision-making, and promoting stewardship to preserve antimicrobial efficacy. Continued investment in laboratory infrastructure, data analysis capacity, and research innovation will be essential to meet the growing challenges of antimicrobial resistance.